I am very happy to announce that my new manuscript has been accepted for publication in arXiv.
https://arxiv.org/html/2604.01475v1
This is my first independent work and the first of a series of works to be published on EEG analysis and neurodegeneration. The work combines physiology, signal analysis & applied mathematics and deep learning techniques.
Interpretable Electrophysiological Features of Resting-State EEG Capture Cortical Network Dynamics in Parkinson’s Disease
Antonios G. Dougalis1,2*
1Computational Neuroscience, Institute of Applied and Computational Mathematics, Foundation for Research and Technology – Hellas (FORTH), 70013, Heraklion, Greece
2Independent Researcher, 50131, Lefkopigi, Greece
* Correspondence, antoniosdougalis@med.uoc.gr; antoniosdougalis@gmail.com, orcID: 0000-0002-2139-1616
Abstract
Background/Objectives: Parkinson’s disease (PD) alters cortical neural dynamics, yet reliable non-invasive electrophysiological biomarkers remain elusive. Resting-state electroencephalography (EEG) can capture complementary spectral, connectivity, and dynamical properties reflecting disease-related neural alterations and medication effects. This study examined whether interpretable EEG features capturing complementary aspects of neural dynamics can discriminate Parkinsonian neural states and provide insight into underlying electrophysiological differences.
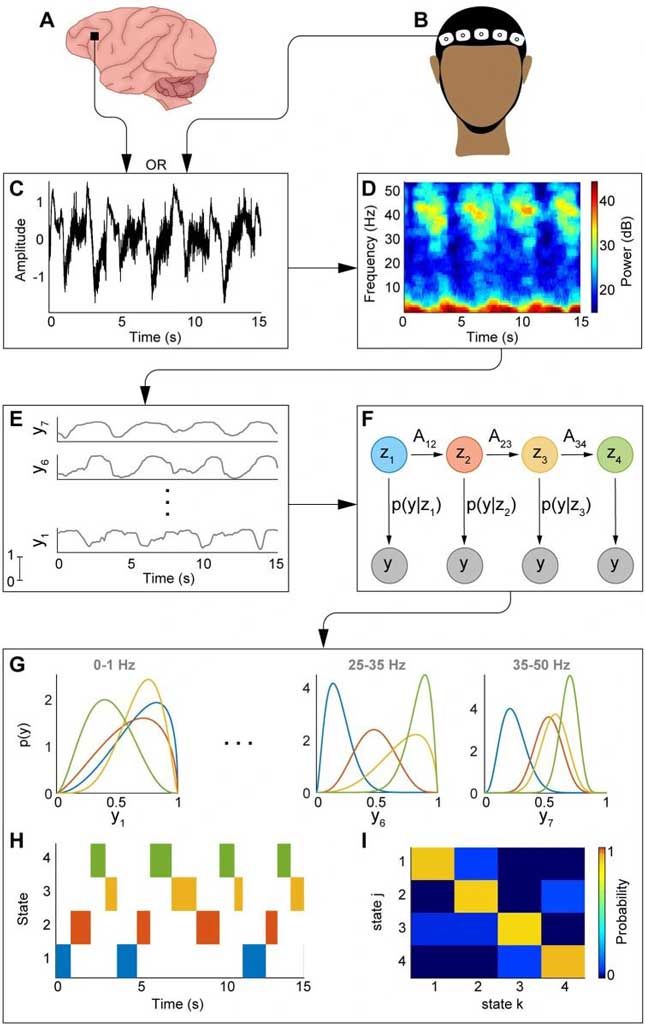
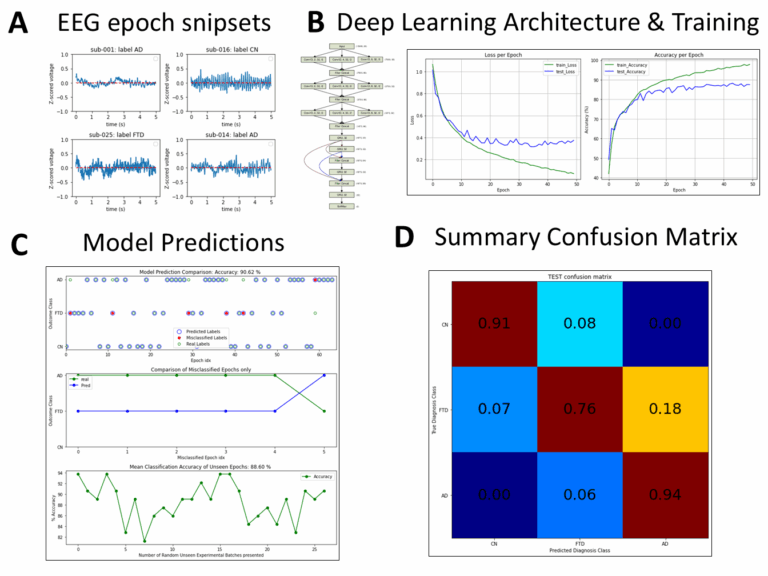
Methods: Resting-state EEG recordings from healthy controls and PD patients recorded in both off- and on-medication states were analyzed. A comprehensive set of interpretable features was extracted from each electrode and grouped into Standard descriptors (spectral power, phase synchronization, time-domain statistics) and Dynamical descriptors (aperiodic activity, cross-frequency coupling, scale-free dynamics, neuronal avalanche statistics, and instantaneous frequency measures). A multi-head attention transformer classifier was trained using strict leave-one-subject-out validation. Feature complementarity was assessed using random ablation analyses and pairwise correlation measures. Group-level comparisons were performed to identify electrophysiological differences associated with disease and medication state.
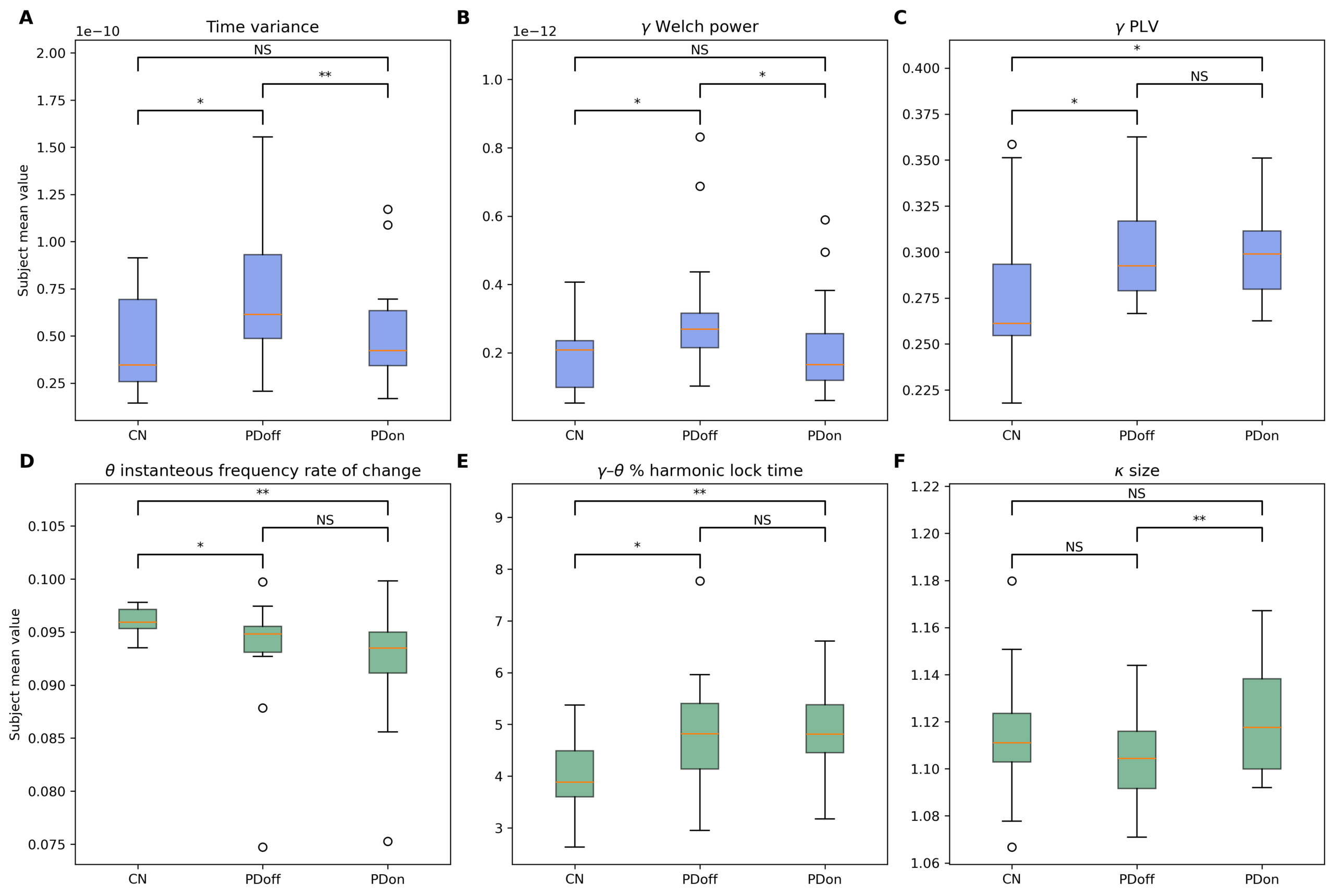
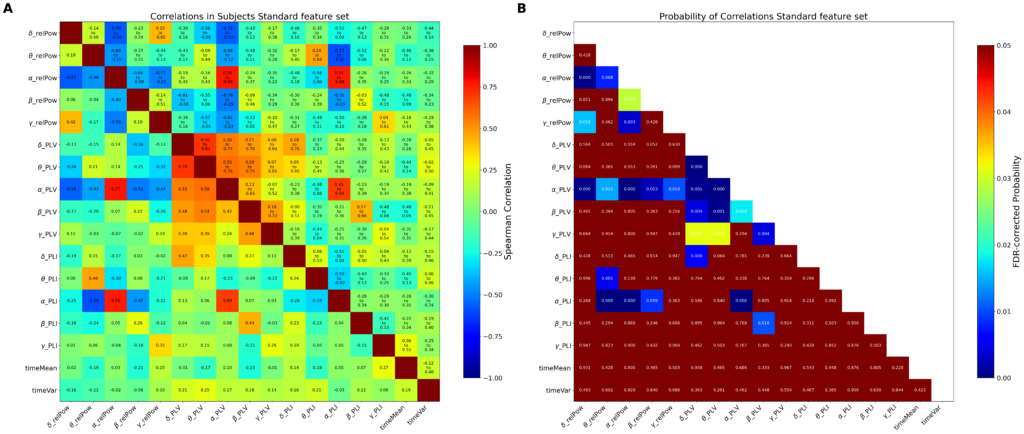
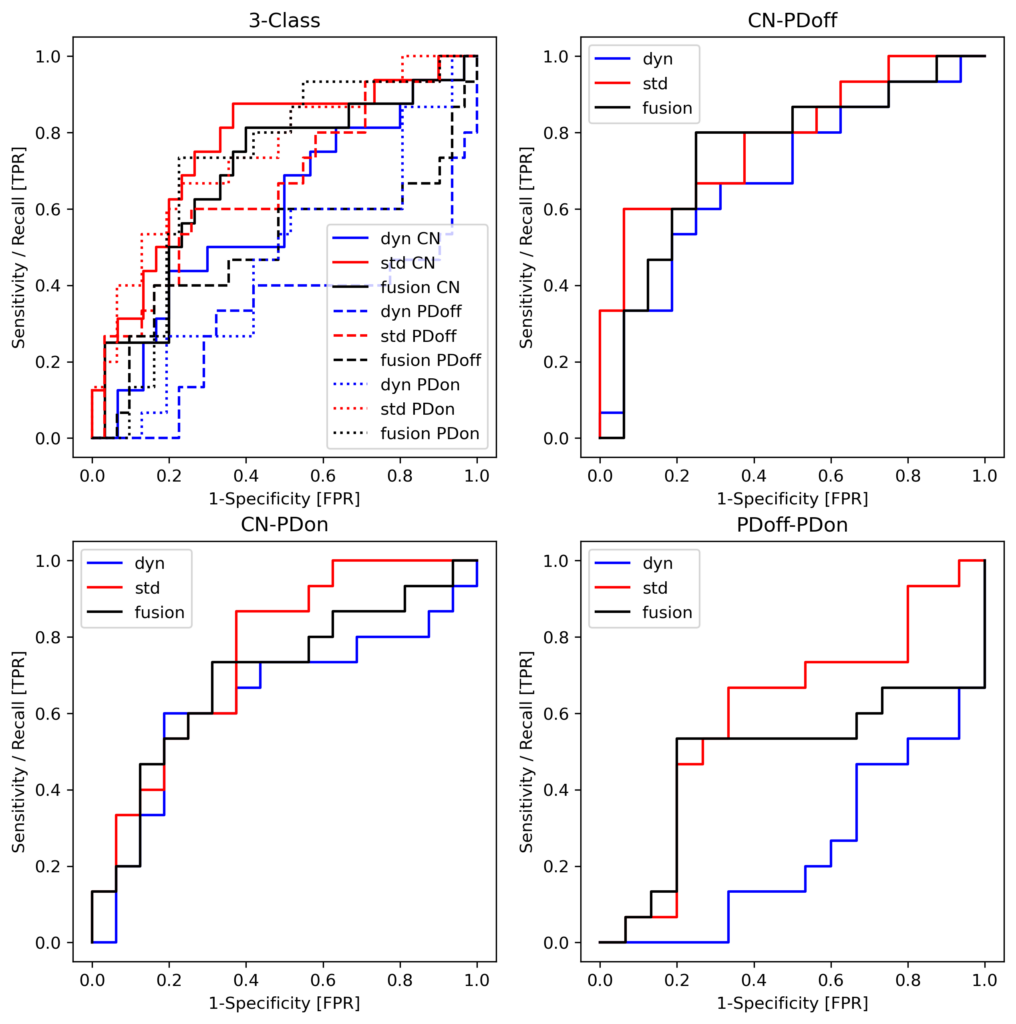
Results: Standard and Dynamical feature sets achieved comparable classification performance. Standard features showed strongest performance in discriminating medication states (PDoff vs PDon), whereas Dynamical features performed competitively in contrasts between PD patients and healthy controls. Random feature ablation analyses indicated that Dynamical descriptors provide complementary information distributed across features while correlation analysis revealed low redundancy within both feature sets. Group-level comparisons revealed medication-sensitive reductions in delta power and voltage variance, modulation of neuronal avalanche statistics, persistent increases in theta phase synchronization in PD patients, and disease-related alterations in cross-frequency interactions.
Conclusions: Distributed electrophysiological descriptors derived from resting-state EEG capture complementary aspects of Parkinsonian neural dynamics. Traditional spectral and synchronization features primarily reflect medication-related neural modulation, whereas dynamical descriptors reveal broader alterations in cortical network organization associated with disease but also with medication. These findings support multivariate EEG representations as a promising framework for developing non-invasive biomarkers of Parkinson’s disease.
Follow this space for more in depth discussion on the subject