#EEG Neurobiomarkers via Machine learning #machinelearning analysis in #neuroscience #dataanalysis, #sleep
I have recently been talking about machine learning tools and applications in neuroscience in a rather cryptic manner.
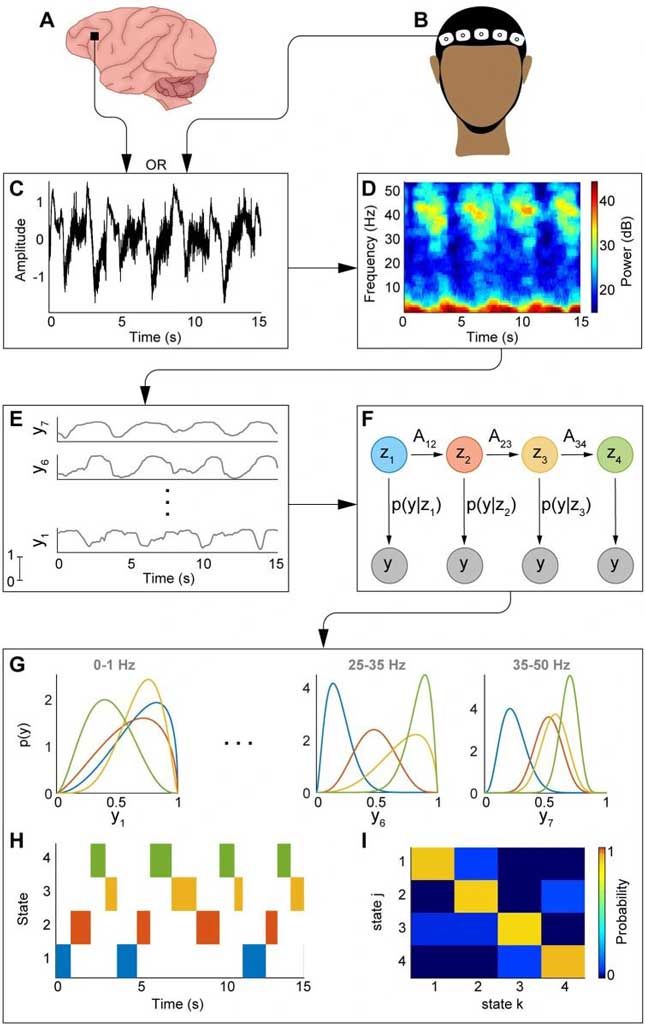
To give you some context over my interests in neuroscience and modern #ML tools, below I attach one of my favourite manuscripts that applies machine learning & statistical tools for state- feature classification of EEG data from humans under ketamine anaesthesia using a five state hidden Markov model (HMM).
https://lnkd.in/eWD7jtTi
Garwood IC, Chakravarty S, Donoghue J, Mahnke M, Kahali P, Chamadia S, Akeju O, Miller EK, Brown EN. A hidden Markov model reliably characterizes ketamine-induced spectral dynamics in macaque local field potentials and human electroencephalograms. PLoS Comput Biol. 2021 Aug 18;17(8):e1009280. doi: 10.1371/journal.pcbi.1009280. PMID: 34407069; PMCID: PMC8405019.
The code for their routines, that is written exclusively in MatLab, can be found here:
https://lnkd.in/eC7bQXeJ
Algorithms like these presented can classify features from long stretches of EEG data with high reliability and can be used to address issues of sleep-wakefulness architecture and link those to underlying pathology given clinical data are also available.
Ultimately, deriving neurobiomarkers as EEG features from specific disease, genetic or neurodevelopmental status could be possible.
Just think of the potential this has in disease diagnosis, classification, severity scoring and progression.
Despite the modernity of these tools the indispensable reading material for anyone interested in the applications in this field comprises the two following works:
Machine Learning Toolbox, Version 1.00, Zoubin Ghahramani, University of
Toronto (1996)
Hidden Markov Model (HMM) Toolbox, Kevin Murphy (1998)
EEG BIOMARKERS VIA MACHINE LEARNING ANALYSIS